Haritica
Bioinformatics Without Code
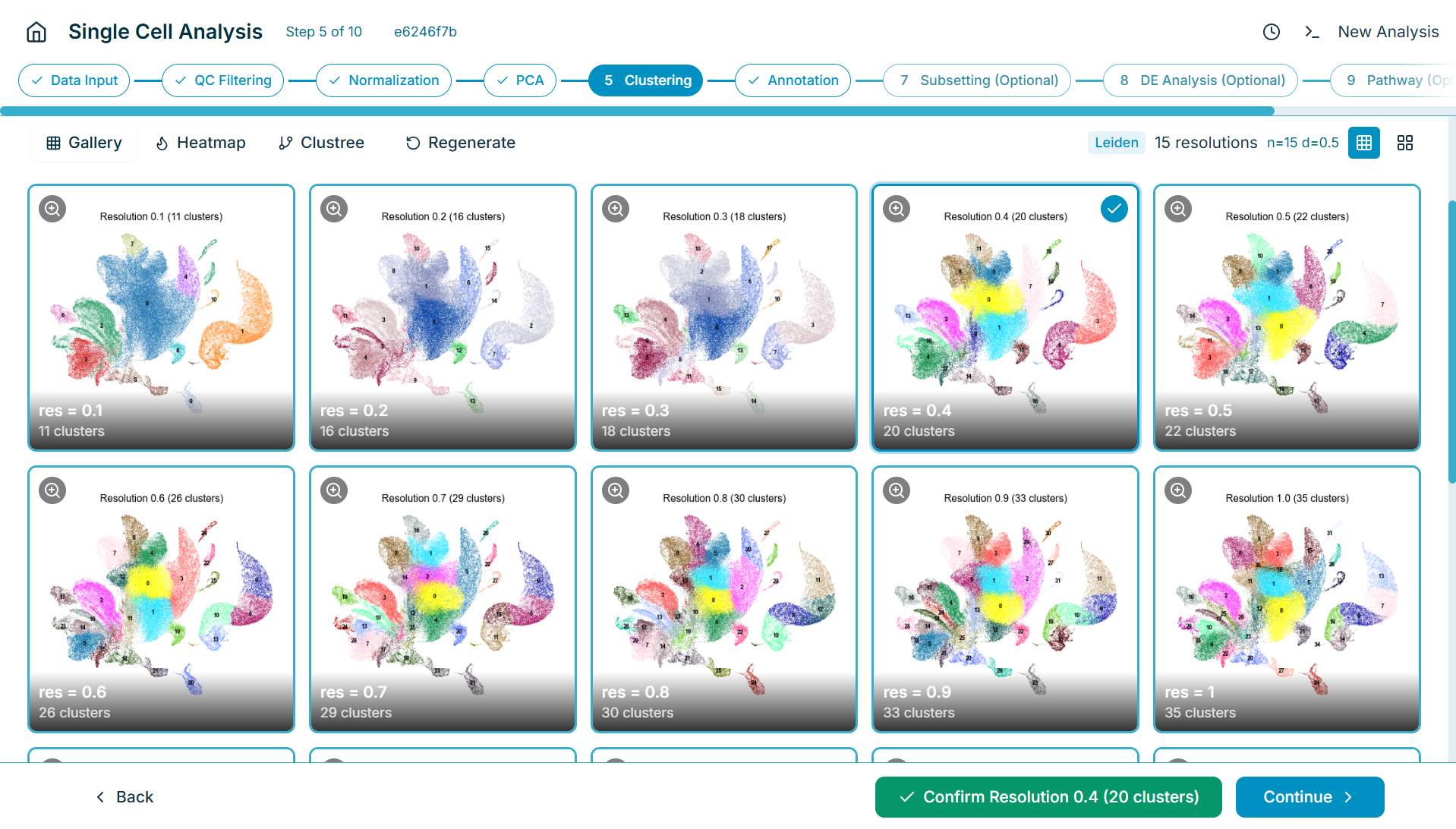
Single-Cell UMAP with Cluster Annotation
Bioinformatics Without Code

Single-Cell UMAP with Cluster Annotation
From raw sequencing reads to publication-ready figures.
Step 1
Select your raw sequencing files from any format — FASTQ, BAM, or CSV count matrices.
Step 2
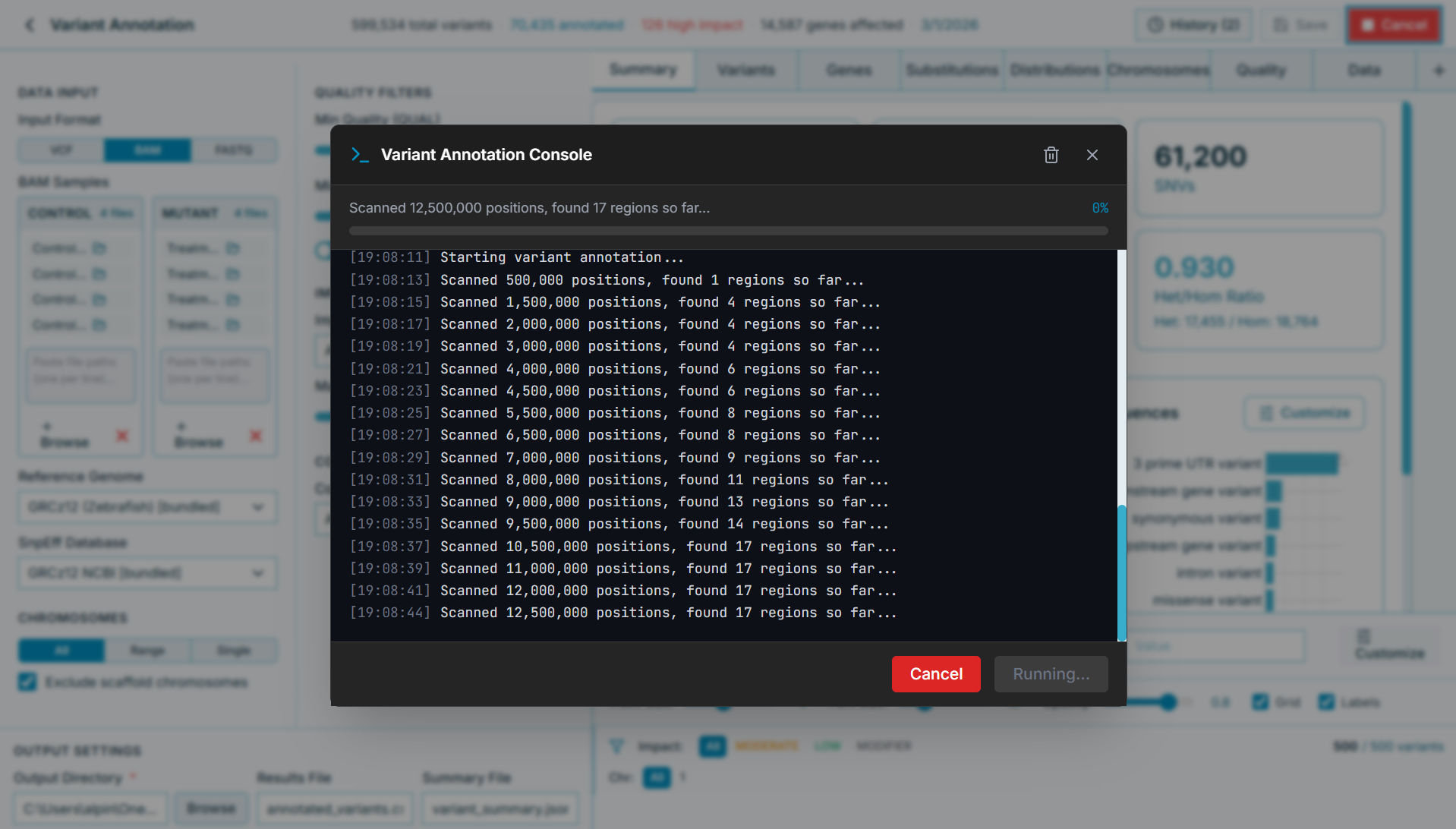
Bundled HISAT2 aligns your reads automatically. Watch progress in real time — no terminal needed.
Step 3
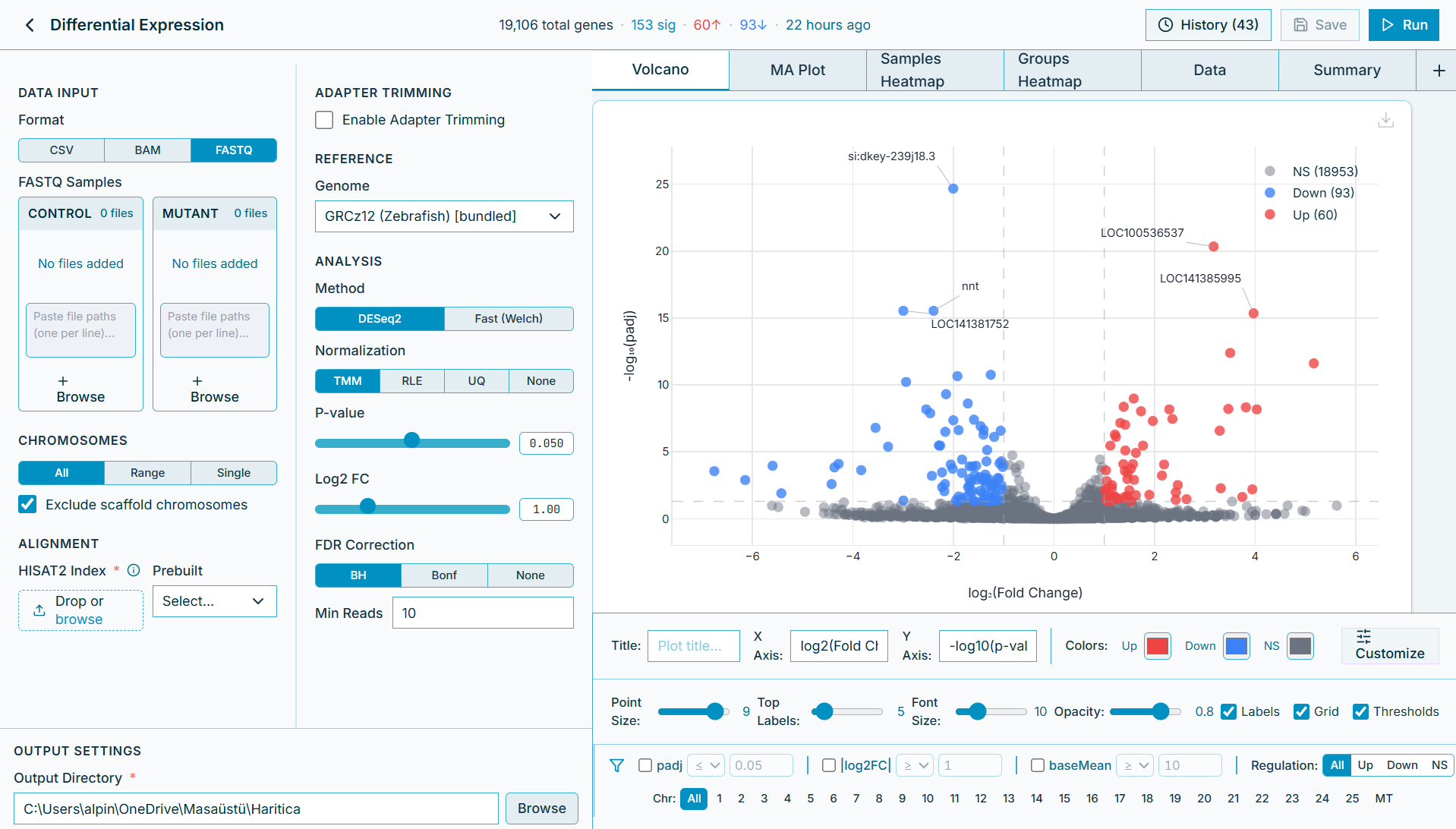
Interactive volcano plots, heatmaps, and tables ready for your next paper. Customize every detail.
The old way
At least 7 commands. At least 3 tools. At least 1 hour just to prepare files.
With Haritica
Drop FASTQ files here
or click to browse
Drag. Drop. Done.
No conda. No module load. No PATH. Everything ships with the app.
Select your question. Haritica recommends the right statistical method.
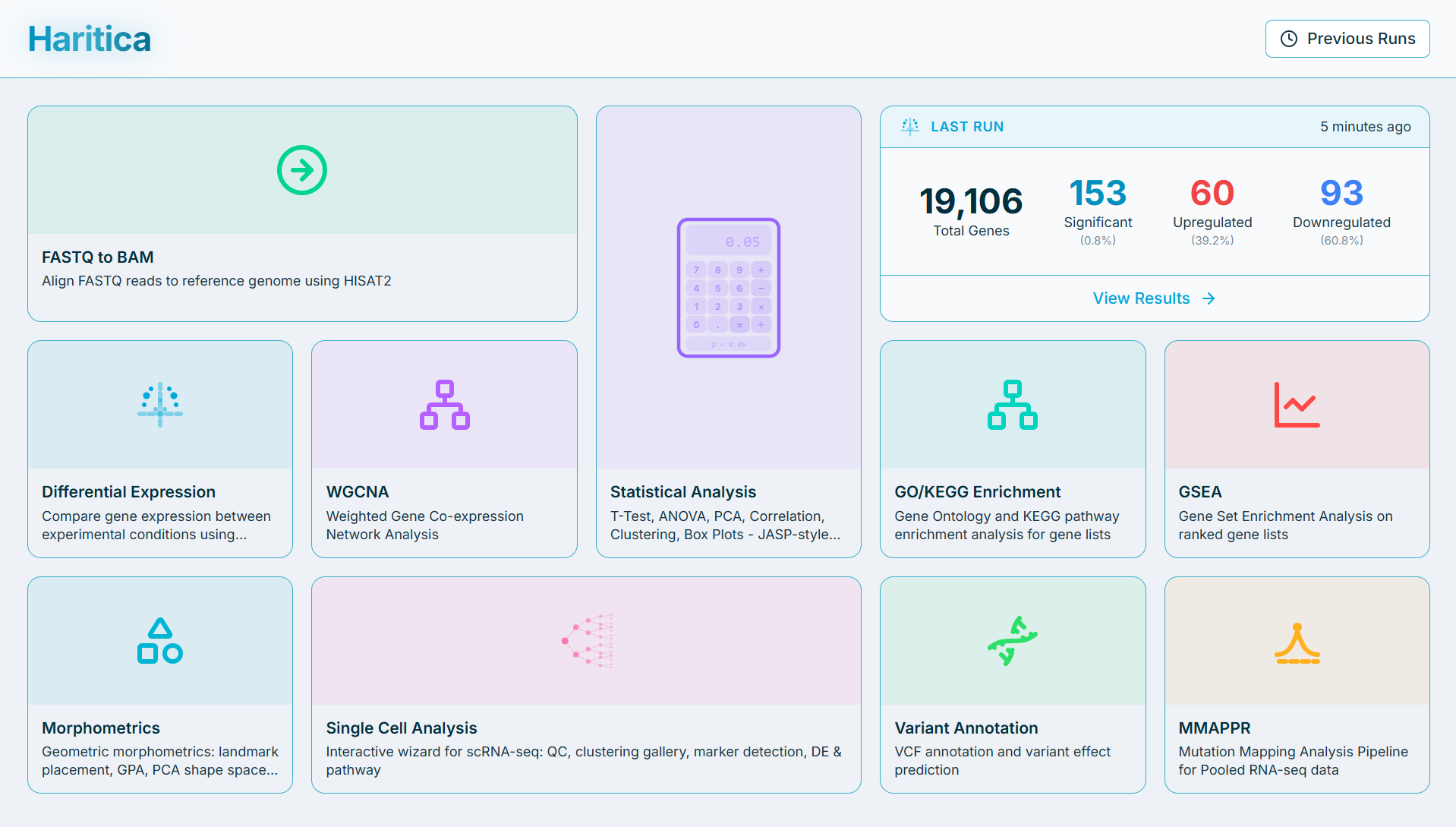
WGCNA network modules, single-cell UMAP clustering, volcano plots, enrichment pathways, and more.
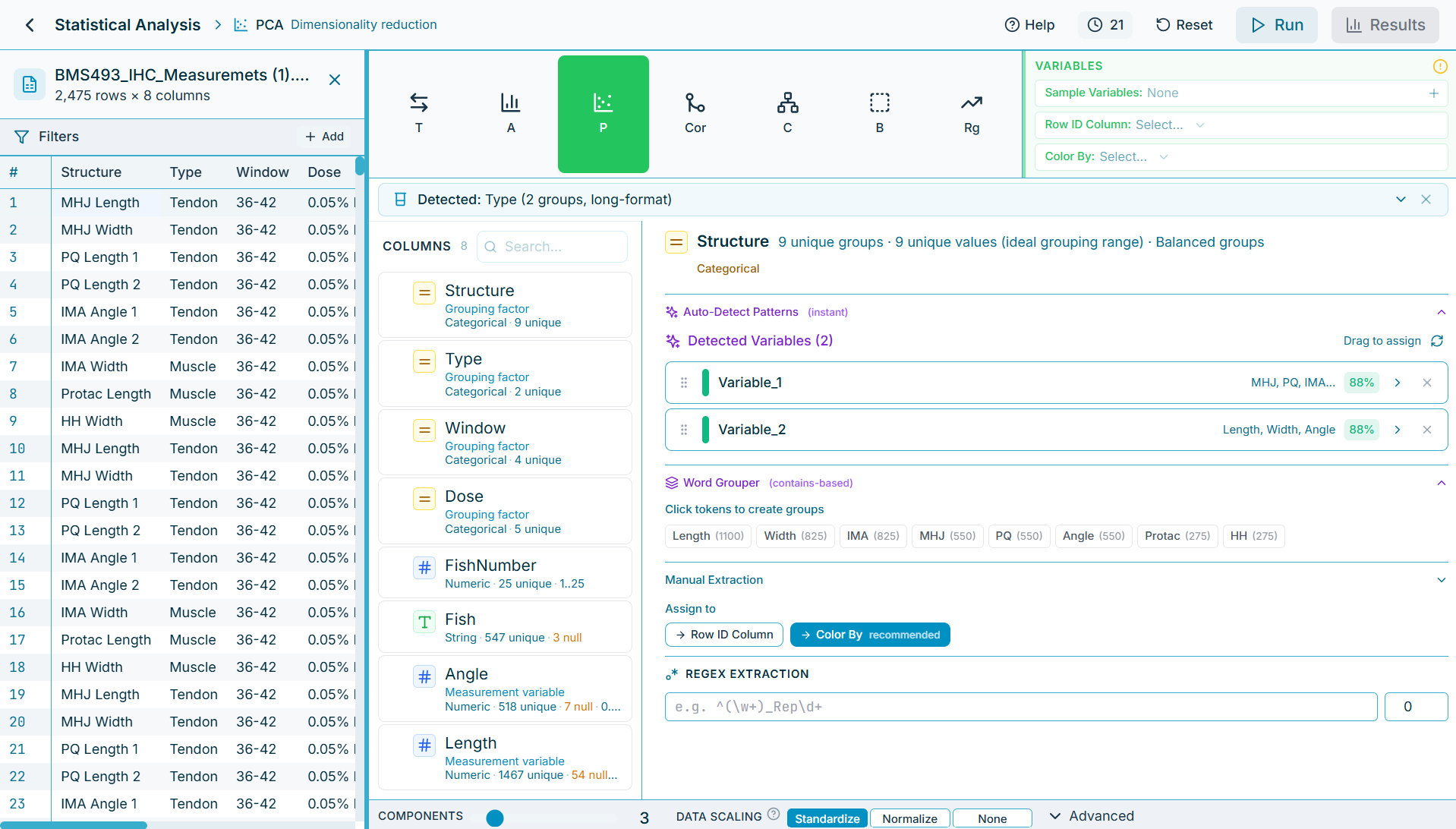
Auto-detection of variables, even if multiple per column

Single-Cell UMAP with Cluster Annotation
The desktop app processes your data on your machine. Cloud compute is available when jobs need to scale beyond your local resources.